DACTRL: Thalamic PGES Detection
Bhargava Ganthi · PhD Research · 26+ Experiments · April 2026
Problem & Goal
PGES (Post-Ictal Generalized EEG Suppression) is the strongest known electrographic risk marker for SUDEP (Sudden Unexpected Death in Epilepsy). Longer PGES duration directly predicts higher SUDEP risk. A sensing-enabled DBS device (Medtronic Percept PC) can trigger alerts automatically — but no public thalamic PGES dataset exists, and only 15 patients were available. Standard deep learning is infeasible. The thesis asked: Can few-shot learning bridge the gap, and can large public scalp EEG corpora (TUH) help?
Biological Discovery — Perspective Inversion
Before any ML code, we verified clinical PGES detection rules on thalamic recordings. Applying scalp algorithms naively gave F1=0.400 — worse than random chance.
Feature Direction Inversion Table
| Feature | Scalp PGES | Thalamic PGES | Direction |
|---|---|---|---|
| Suppression Ratio | HIGH (flat signal) | LOW (active delta) | ⚠️ INVERTED |
| RMS Amplitude | LOW | HIGH | ⚠️ INVERTED |
| Zero Crossings | LOW | HIGH | ⚠️ INVERTED |
| Approx Entropy | LOW | LOW | ✓ Same |
| Spectral Ratio (δ/α) | HIGH | HIGH | ✓ Same |
| Shannon Entropy | LOW | LOW | ✓ Same |
Confirming the Hypothesis — Simultaneous Paired Recordings
To verify that scalp and thalamic signals truly encode the same PGES event from opposite perspectives, we identified 3 patients (P2, P10, P12) with adequate simultaneous scalp + thalamic coverage during seizures. We trained a shared encoder on the same seizure from both recording sites simultaneously — forcing the model to bridge the two perspectives within a single embedding space.
DACTRL-TSM Architecture
A 4-layer Causal Transformer pre-trained self-supervisedly on 8-window sequences (40s context) via next-window cosine+MSE prediction. No labels required for pre-training. At test time, K labeled windows seed a ProtoNet classifier.
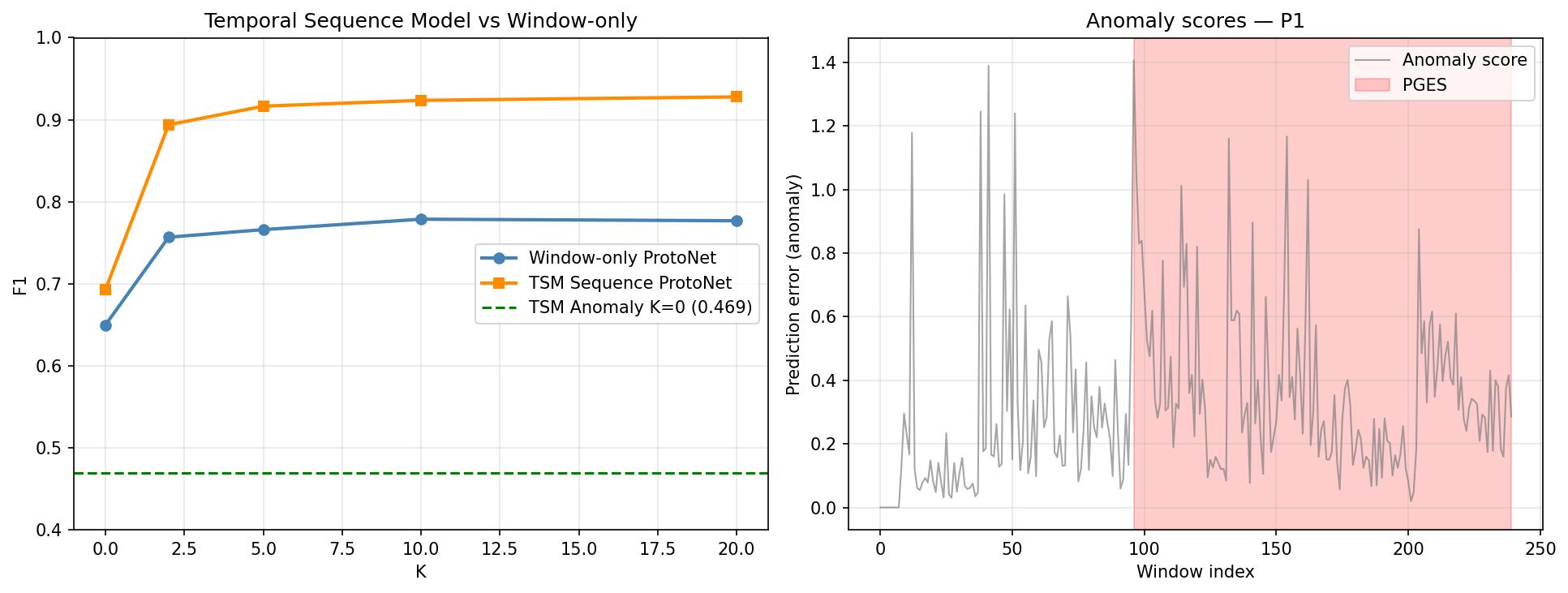
Why Temporal Context? The Core Motivation
How Data is Structured for TSM
Each patient recording is split into 5-second windows. For each window we compute the 17 features above, yielding a 17-dimensional vector per window. These vectors are then grouped into sequences of 8 consecutive windows (N_CTX=8), creating a sequence of shape [8 × 17]. Each sequence therefore covers exactly 40 seconds of continuous thalamic LFP.
| Stage | What happens | Why |
|---|---|---|
| Raw EDF → 5s windows | Each LFP recording is segmented into 5-second non-overlapping windows | 5s is long enough to estimate spectral features reliably; short enough for temporal resolution |
| 17 features per window | Each window → 17-dim feature vector (time-domain + spectral + complexity) | Captures different aspects of the signal; robust to amplitude noise; interpretable |
| Group into 8-window sequences | N_CTX=8 consecutive windows = 40s sequences | 40s context covers a full PGES onset pattern; ablation shows this is optimal (N_CTX={4..16} within ±0.007 F1) |
| StandardScaler on train only | Fit scaler on N−1 training patients; apply to test patient without refitting | Prevents data leakage; each patient contributes to normalization only as a training patient |
| Pre-training (self-supervised) | CausalTransformer predicts window t+1 features from windows 1..t | No labels needed; model learns what "normal thalamic dynamics" look like across time |
| Test-time ProtoNet (K-shot) | K labeled sequences → two prototype vectors (PGES, baseline); new sequence classified by cosine distance to nearest prototype | Few-shot: K can be as low as 2 (one labeled seizure is enough) |
Architecture Details
| Component | Value | Why this choice |
|---|---|---|
| Model type | CausalTransformer (4-layer) | Causal masking enforces real-time constraint; transformer captures long-range dependencies |
| D_MODEL | 64 | Matched to 17-feature input after projection; large enough for representation, small enough for N=14 patients |
| N_HEADS | 4 | 4 heads × 16 dim each; captures multiple attention patterns (onset, sustained state, recovery) |
| N_LAYERS | 4 | Ablated: 2-layer underfits, 6-layer overfits at N=14 |
| N_CTX | 8 windows = 40 seconds | Optimal from ablation across {4,6,8,12,16}; all within ±0.007 F1 — robust choice |
| Window size | 5 seconds | Standard in EEG feature extraction; long enough for spectral estimation, short enough for temporal resolution |
| Pre-training loss | Cosine similarity + MSE (next-window prediction) | Cosine encourages directional alignment; MSE constrains magnitude. Together they enforce both "shape" and "scale" consistency |
| Few-shot classifier | ProtoNet (cosine similarity) | Prototype = mean embedding of K support examples per class; classification by nearest prototype distance. Non-parametric — no extra parameters to overfit |
| Training protocol | LOSO, N=14, StandardScaler on train only | LOSO is the most rigorous evaluation for small N; scaler fitted only on training patients prevents leakage |
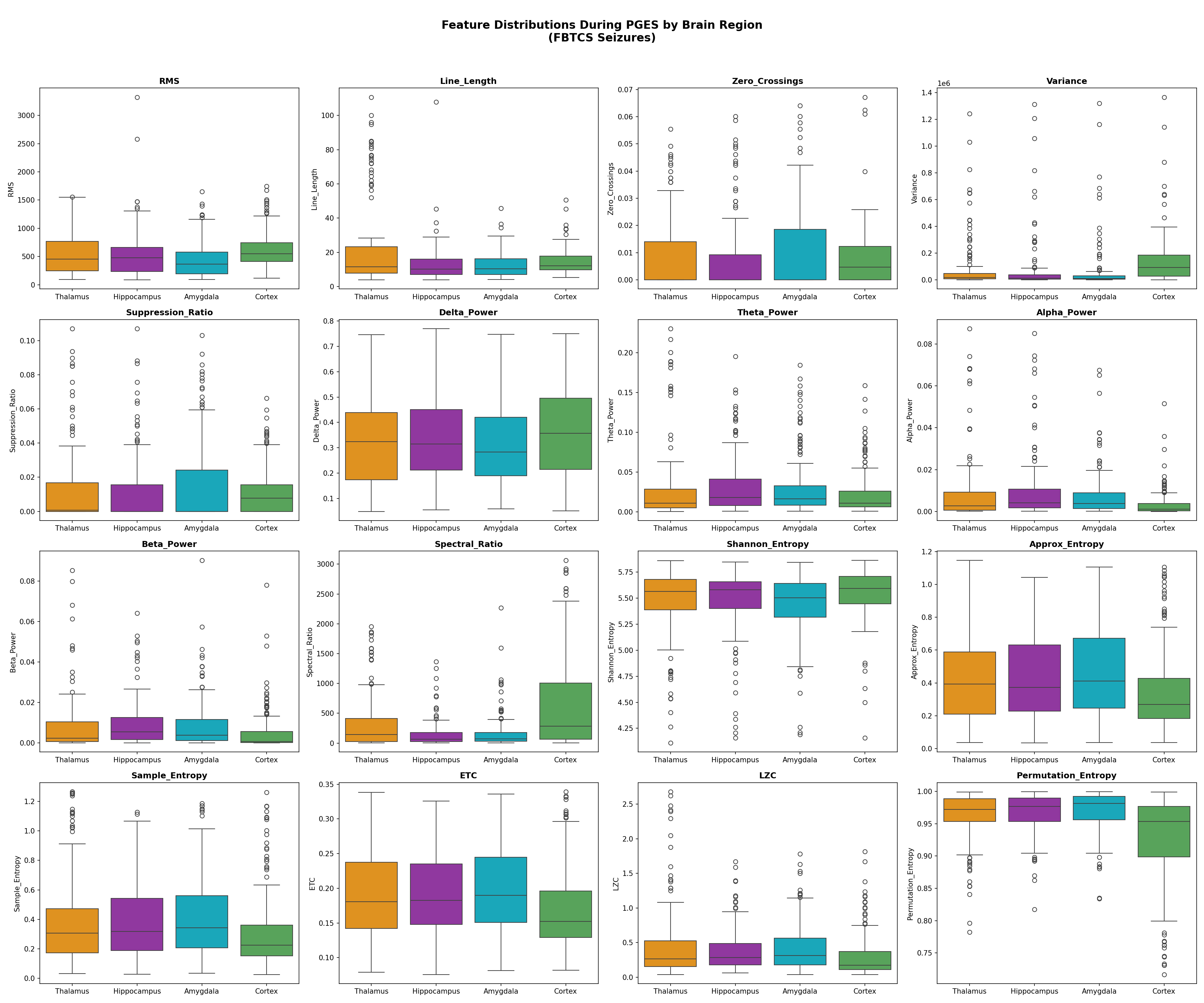
17 Signal Features
Gamma Power was the 17th feature added specifically for thalamic DBS recordings (visible at depth, not on scalp). Features 1–3 carry most discriminative power; features 14–17 contribute marginally but non-negatively.
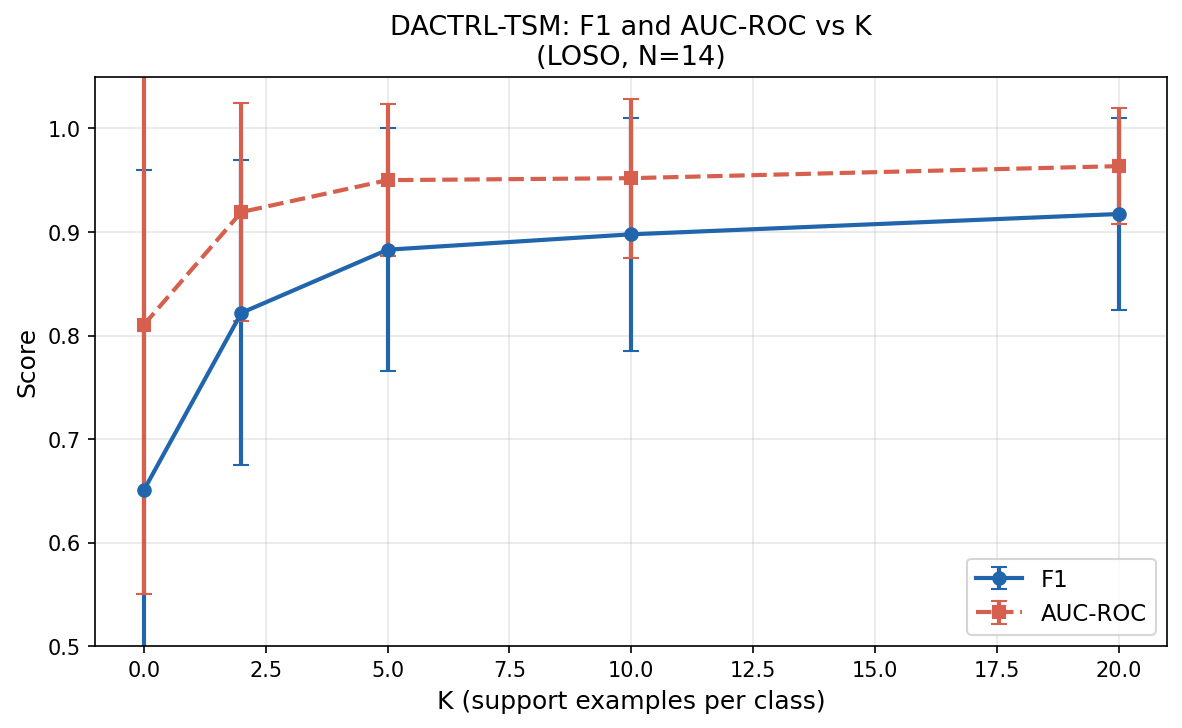
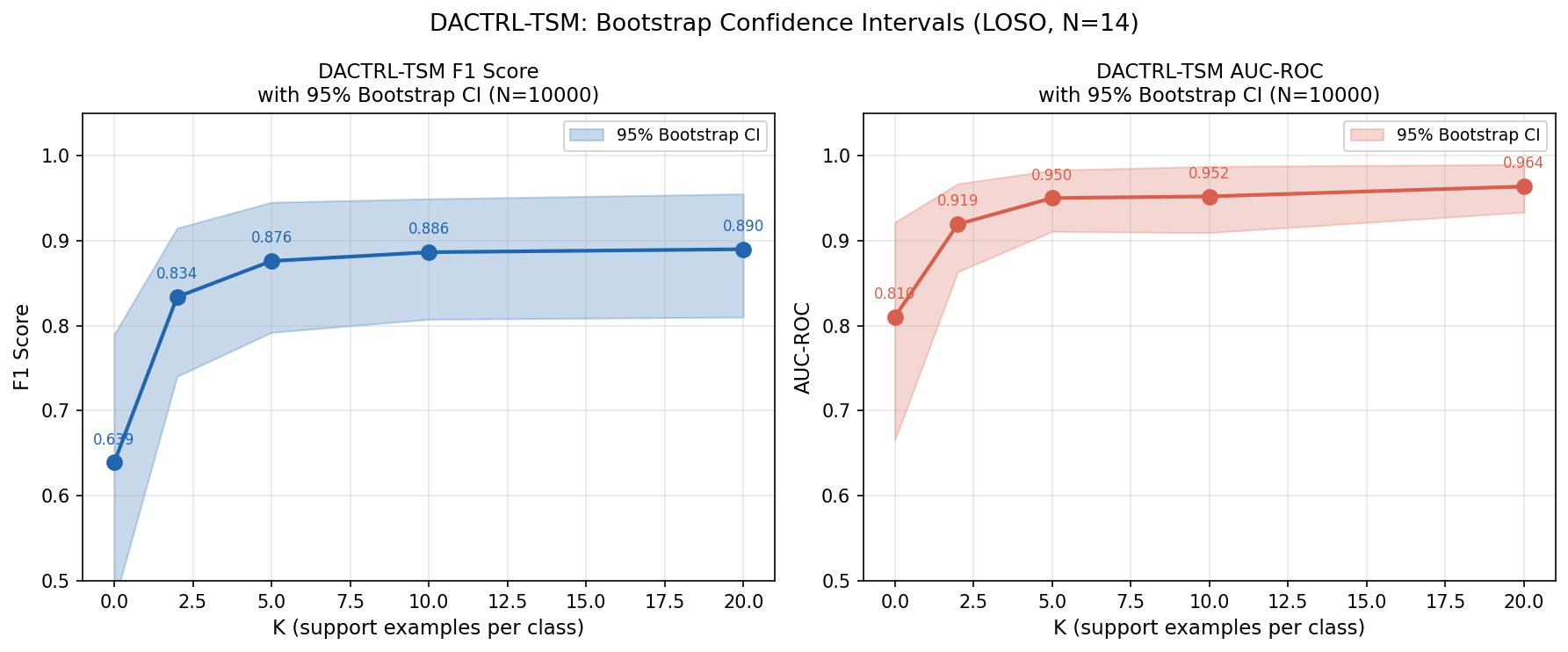
K-Shot Performance
LOSO evaluation on N=14 patients, N_TRIALS=5 averaged. Temporal pre-training was the single largest gain in the project (+24.7pp over zero-shot).
| K | F1 (mean±std) | AUC | 95% Bootstrap CI (F1) |
|---|---|---|---|
| 0 (oracle zero-shot) | 0.640 ± 0.309 | 0.810 | [0.475, 0.790] |
| 2 | 0.834 ± 0.147 | 0.919 | [0.740, 0.915] |
| 5 | 0.876 ± 0.117 | 0.950 | [0.792, 0.945] |
| 10 | 0.898 ± 0.112 | 0.952 | [0.808, 0.949] |
| 20 | 0.917 ± 0.093 | 0.964 | [0.810, 0.955] |
Statistical Significance vs Comparators (Wilcoxon, N=8 confirmed LT/LTP patients)
| Comparator | DACTRL-TSM K=10 | Comparator F1 | ΔF1 | p-value | Sig. |
|---|---|---|---|---|---|
| Zero-shot (K=0) | 0.886 | 0.639 | +0.247 | 0.0009 | ** |
| TSM K=2 | 0.886 | 0.834 | +0.053 | 0.0009 | ** |
| Threshold Rule | 0.886 | 0.696 | +0.190 | 0.004 | ** |
| XGBoost (LOSO) | 0.886 | 0.708 | +0.178 | 0.017 | * |
| Random Forest | 0.886 | 0.715 | +0.171 | 0.017 | * |
| Logistic Regression | 0.886 | 0.686 | +0.201 | 0.004 | ** |
| SVM K=10 | 0.886 | 0.942 | −0.056 | 0.049 | * SVM wins |
| KNN K=10 | 0.886 | 0.900 | −0.014 | ns | — |
Clinical Metrics (K=10)
| Metric | Value | Clinical Interpretation |
|---|---|---|
| Mean FA rate | 67.5 FA/hr | Driven by P12/P15 (atypical ANT morphology) |
| Median FA rate | 30.8 FA/hr | Better estimate — 50% of patients ≤30.8 |
| Patients with 0 FA/hr | 3 of 14 | P11, P2, P4 — perfect specificity |
| Detection latency (mean) | 18.7s | From PGES onset to first correct detection |
| Detection latency (median) | 14.0s | Within first 2–5% of episode duration |
| Detection rate | 100% | All 14 episodes detected across all patients |
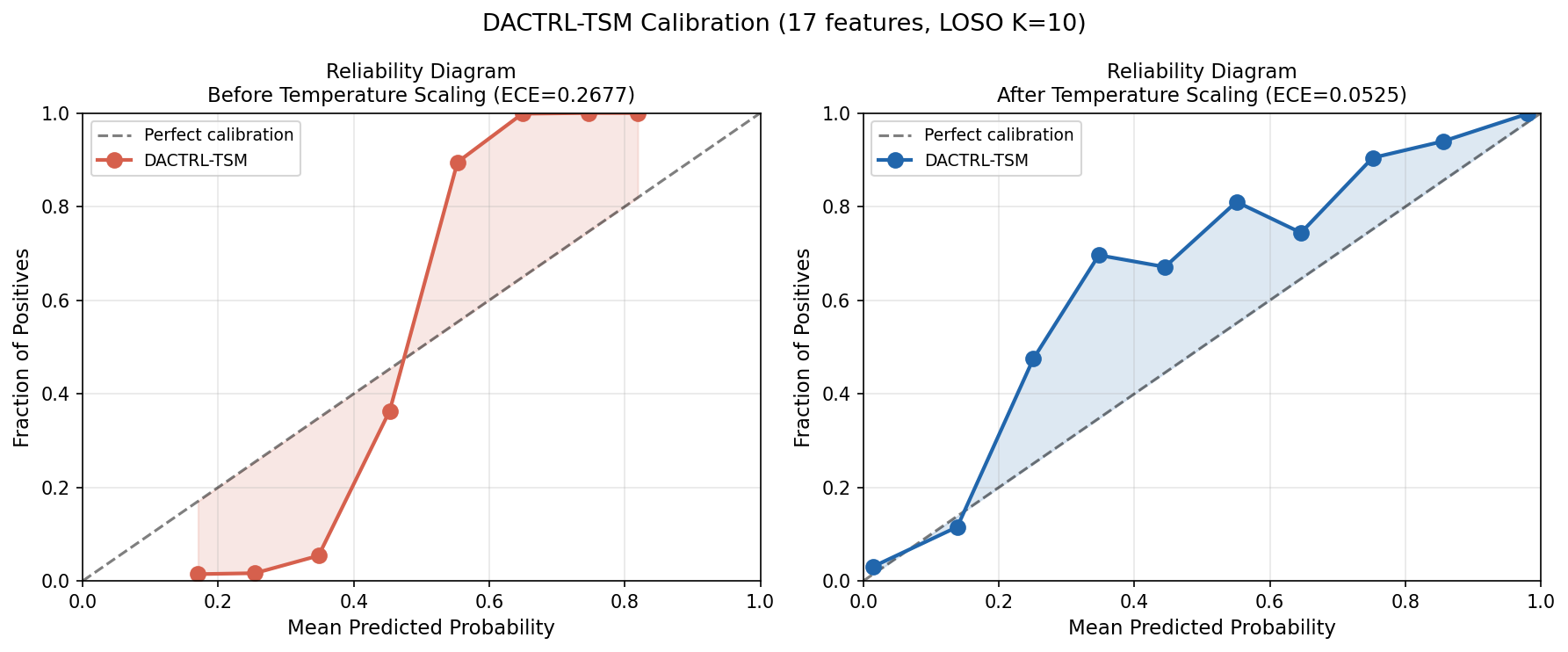
Calibration & Conformal Prediction
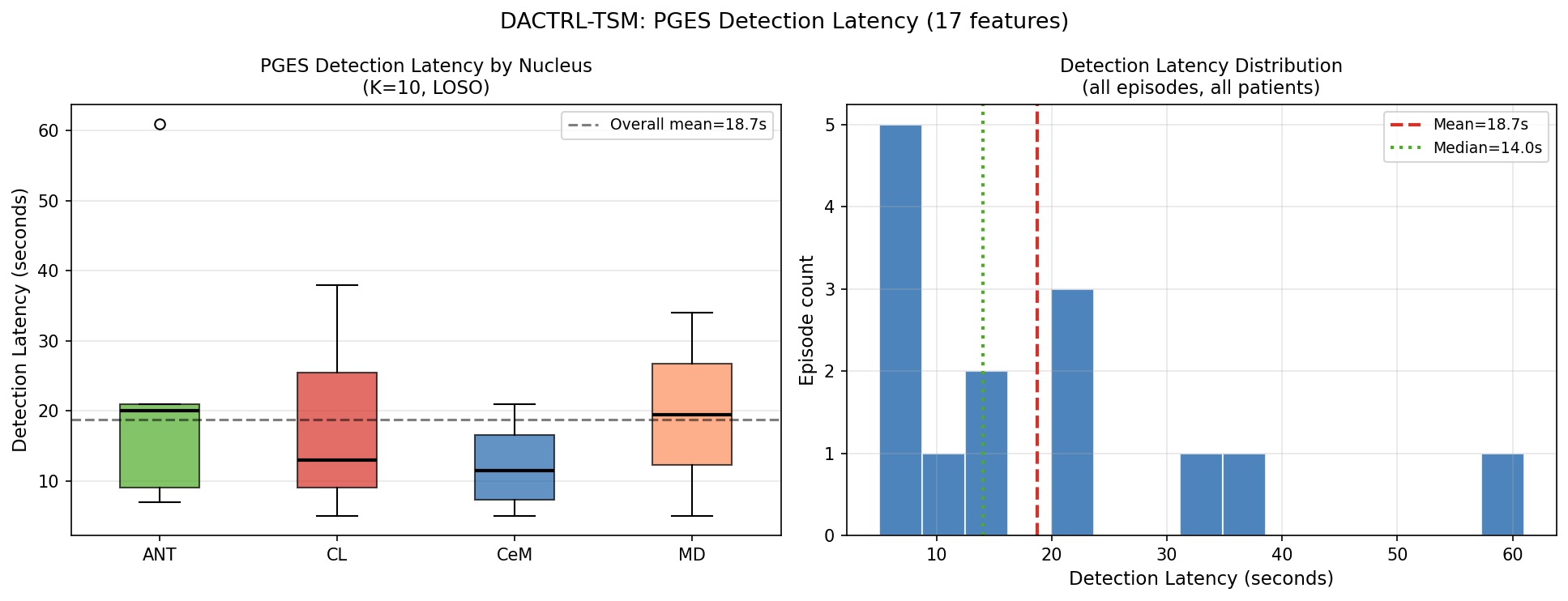
Detection Latency
All 14 PGES episodes detected. Median detection time: 14 seconds from onset.
| Nucleus | Mean (s) | Median (s) | Std (s) | Detection Rate |
|---|---|---|---|---|
| CeM | 12.3 | 11.5 | 7.2 | 100% |
| CL | 18.7 | 13.0 | 17.2 | 100% |
| MD | 19.5 | 19.5 | 20.5 | 100% |
| ANT | 23.6 | 20.0 | 21.8 | 100% |
| Overall | 18.7 | 14.0 | — | 100% |
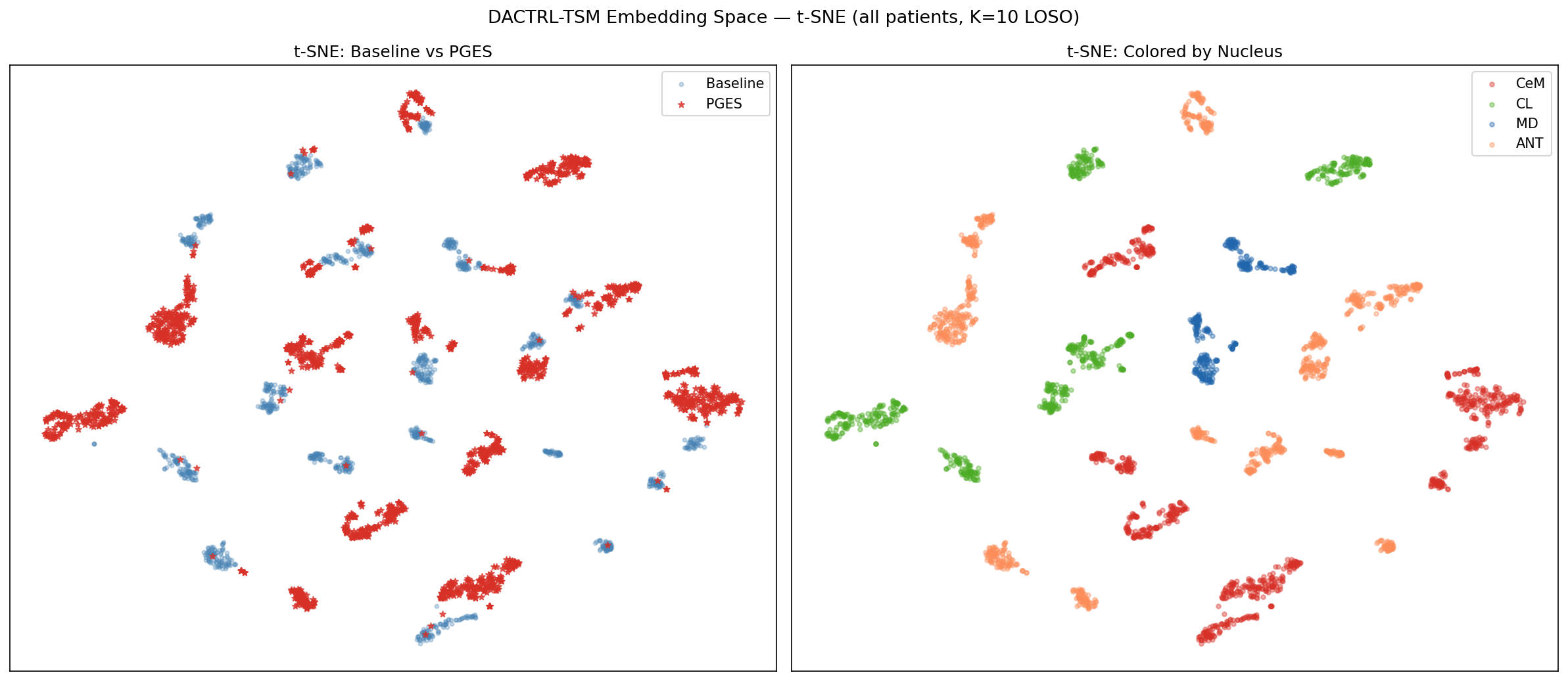
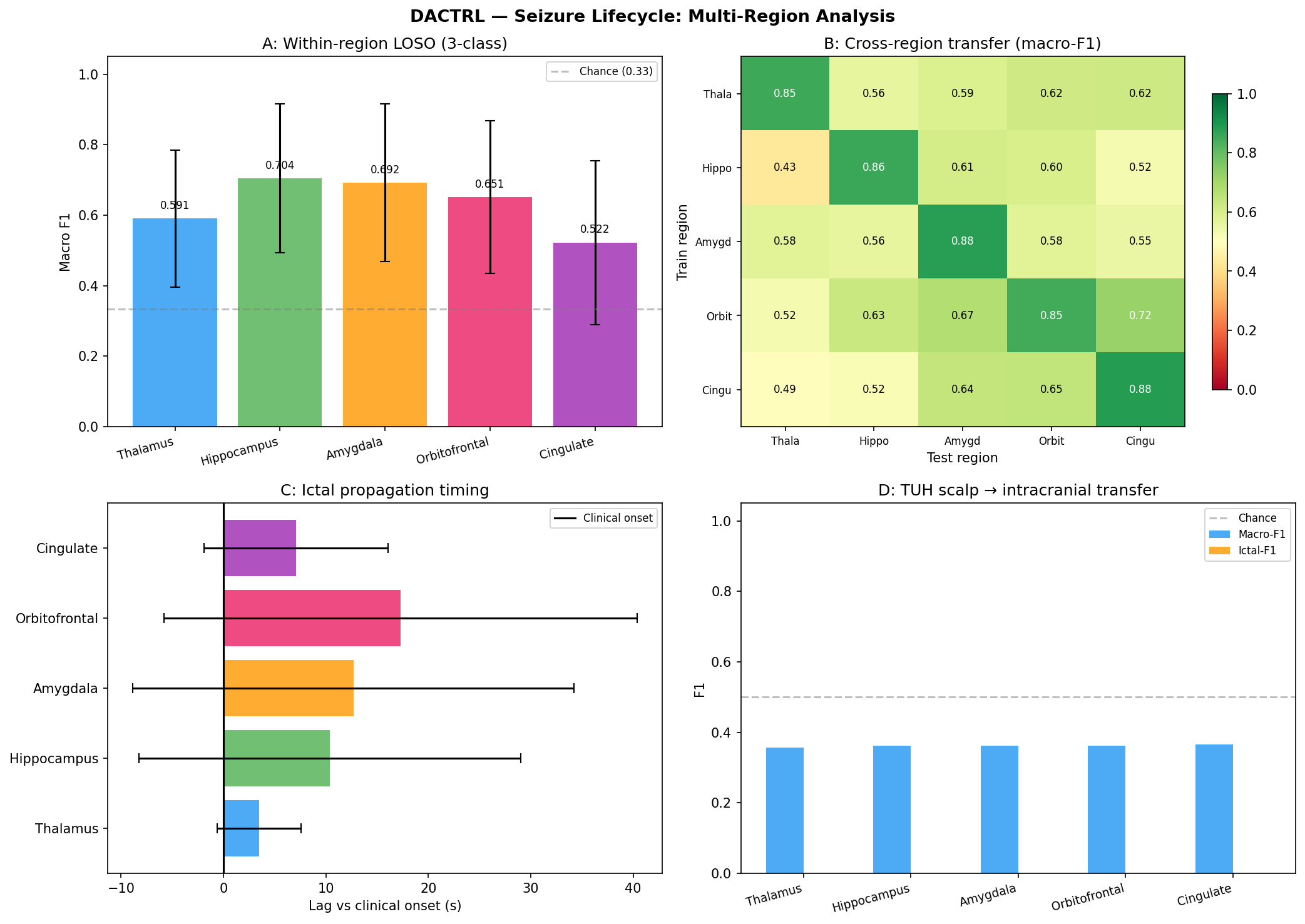
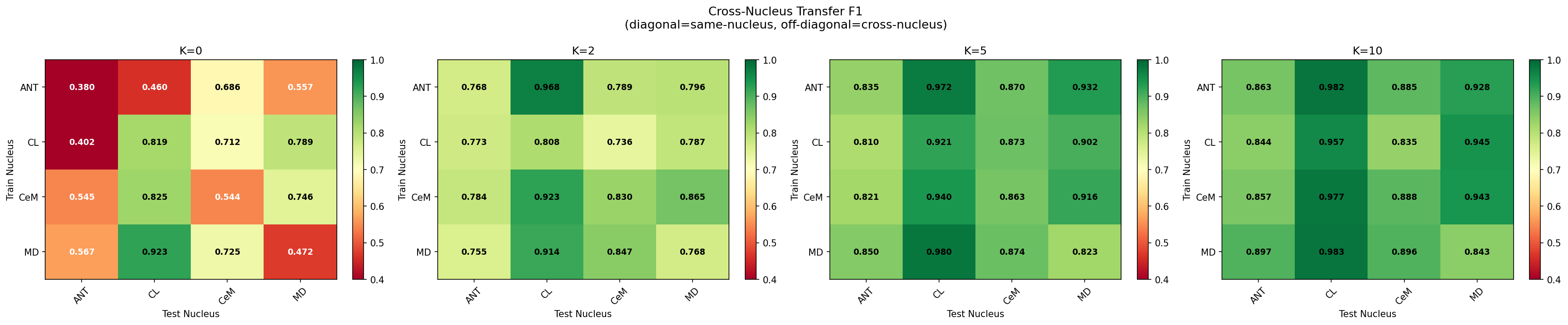
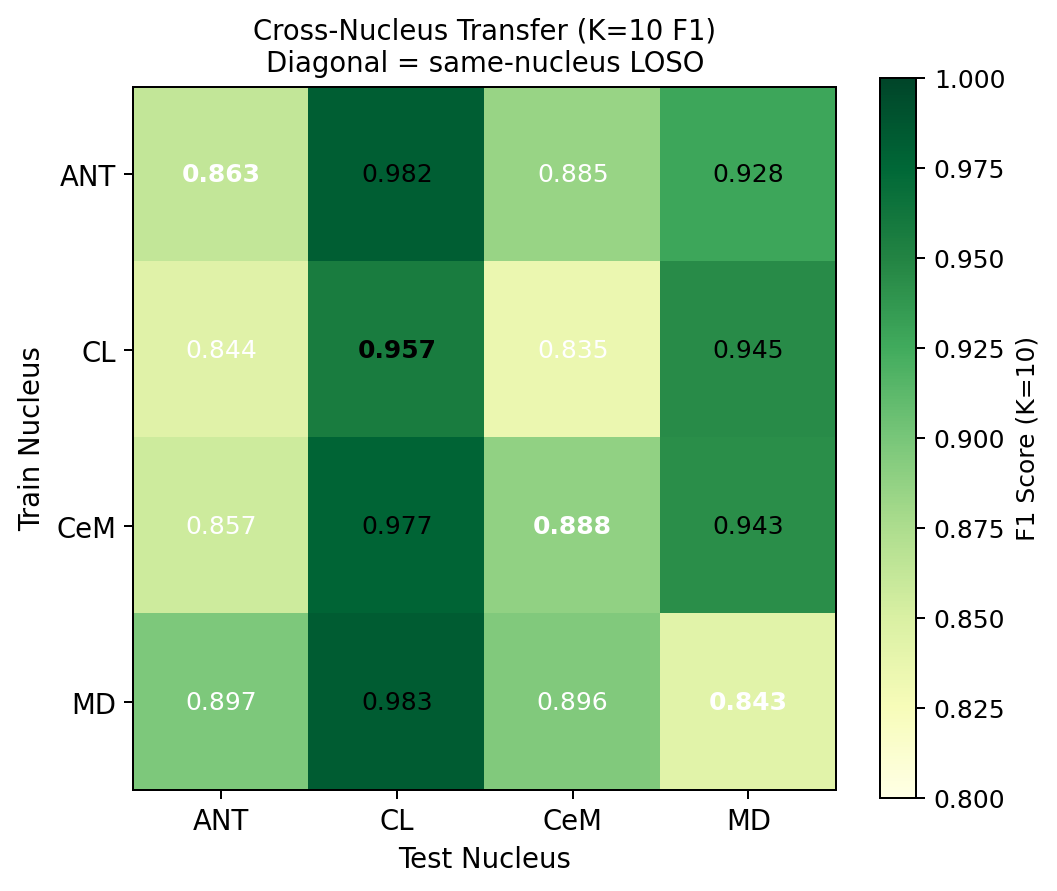
Cross-Nucleus Transfer
Models trained on one nucleus generalise to all others with no degradation. No nucleus-specific models needed.
Scalp Transfer Ablation (12+ Experiments)
Before concluding scalp transfer doesn't work, we exhaustively tested every reasonable approach across 12+ experiments.
| Strategy | K=0 F1 | K=10 F1 | Verdict |
|---|---|---|---|
| Raw scalp encoder | 0.400 | 0.748 | Harmful at K=0 |
| DANN (gradient reversal) | 0.367 | 0.802 | Negative |
| CCA domain mapping | 0.548 | 0.699 | Gap 0.231 vs thalamic |
| TUH-only + thalamic normalization | — | 0.859 | +0.013 (noise) |
| Nucleus-aligned public scalp | — | 0.881 | Best public scalp K>0 |
| Paired encoder (simultaneous records) | 0.747 | 0.793 | Hypothesis confirmed |
| CycleGAN ST_supcon (style transfer) | 0.832 | 0.876 | Best scalp K=0 result |
| Thalamic-only SupCon TSM (B) | 0.678 | 0.913 | Best K≥2 without scalp |
At K≥2: gap collapses to 1.3pp (not statistically significant, p>0.05).
From K=2 onwards, thalamic self-supervised learning alone matches scalp pre-training.
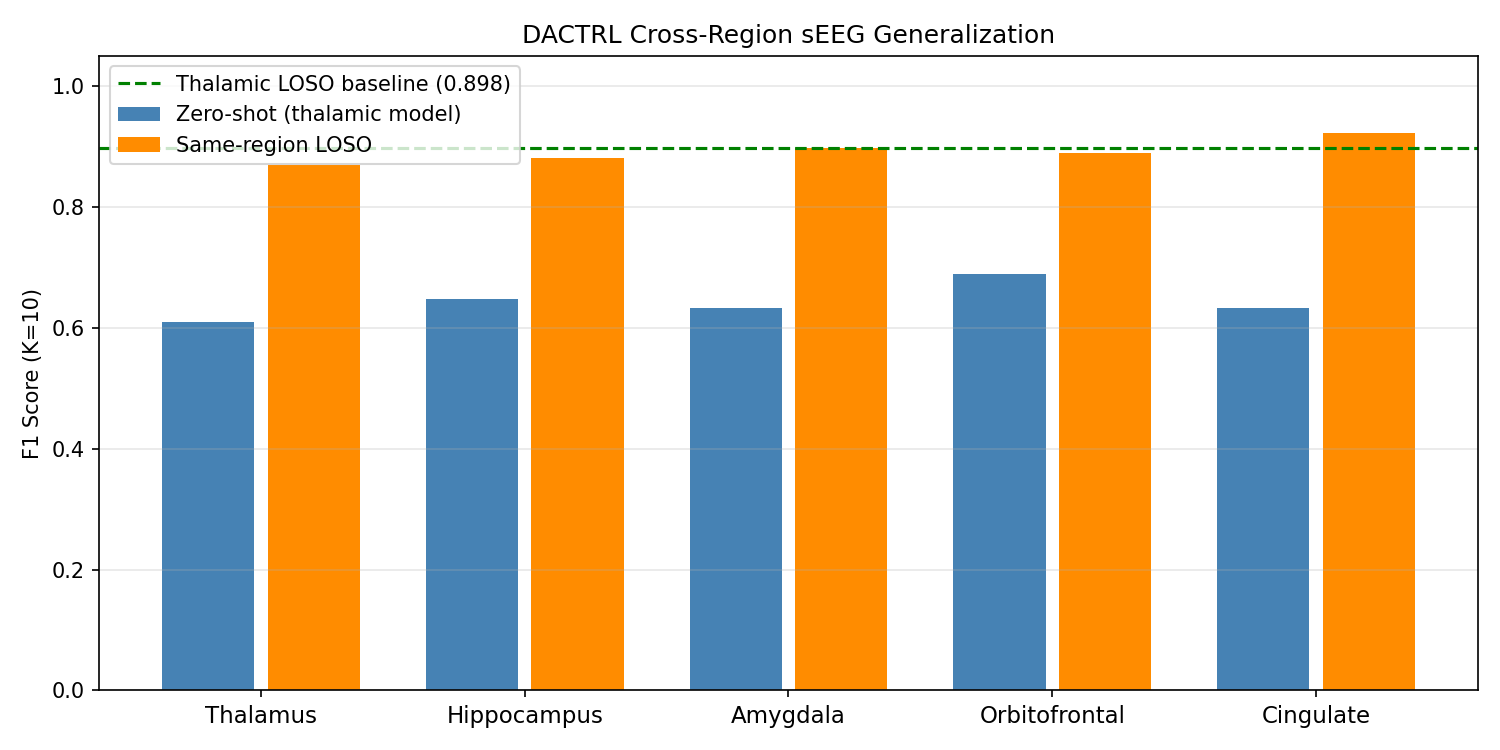
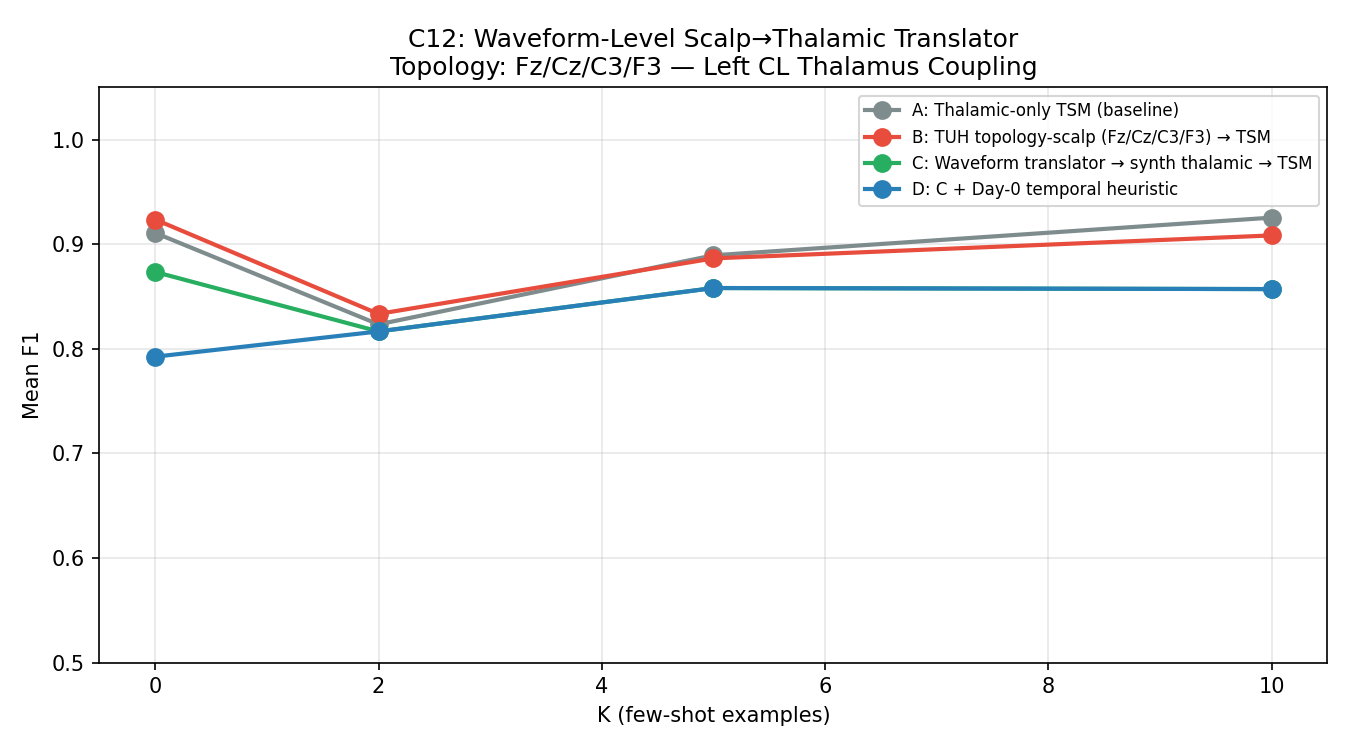
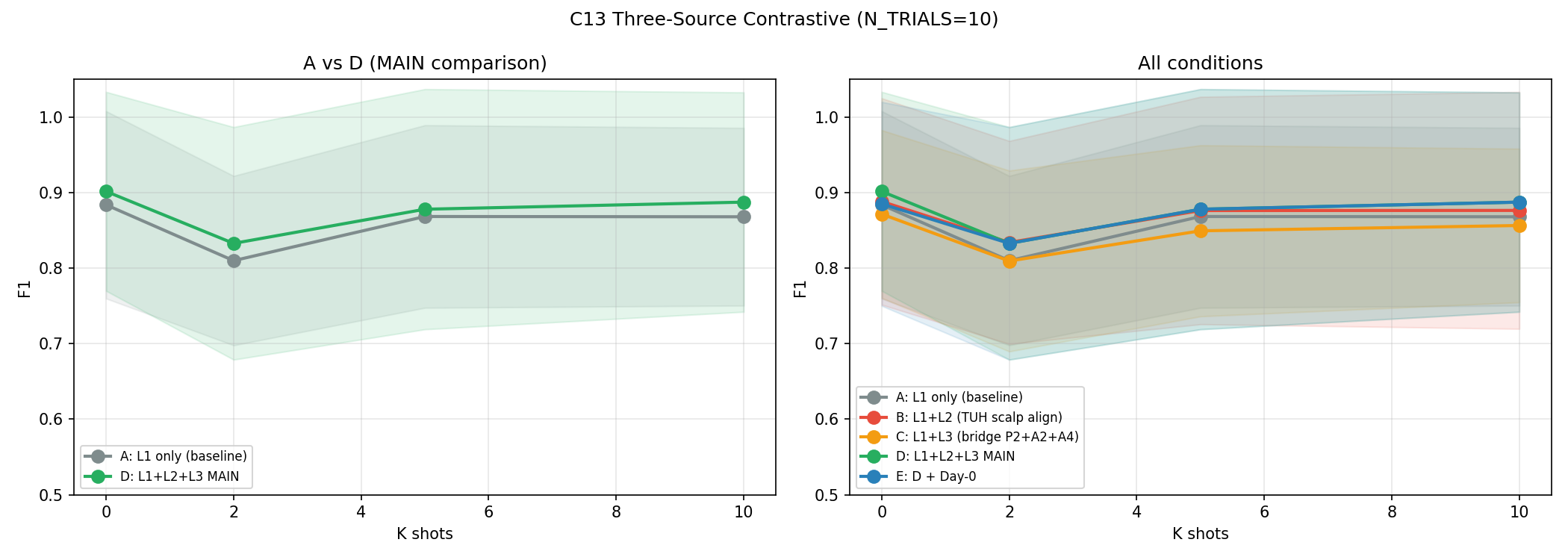
C13 — Three-Source Contrastive (Best Scalp Attempt)
Most sophisticated scalp transfer pipeline: three simultaneous losses (thalamic TSM + TUH↔institutional scalp SupCon + simultaneous scalp↔thalamic bridge). N_TRIALS=10 for statistical power.
| Condition | Description | K=0 F1 | K=2 F1 | K=5 F1 | K=10 F1 |
|---|---|---|---|---|---|
| A | Thalamic TSM only (baseline) | 0.878±0.134 | 0.862±0.130 | 0.867±0.132 | 0.864±0.146 |
| B | +TUH scalp SupCon | 0.869±0.137 | 0.855±0.140 | 0.863±0.141 | 0.860±0.155 |
| C | +Bridge loss | 0.884±0.138 | 0.873±0.128 | 0.879±0.134 | 0.878±0.145 |
| D | +All three losses | 0.901±0.132 | 0.879±0.133 | 0.884±0.130 | 0.887±0.145 |
| E | +ProtoAug | 0.895±0.141 | 0.870±0.139 | 0.877±0.141 | 0.878±0.140 |
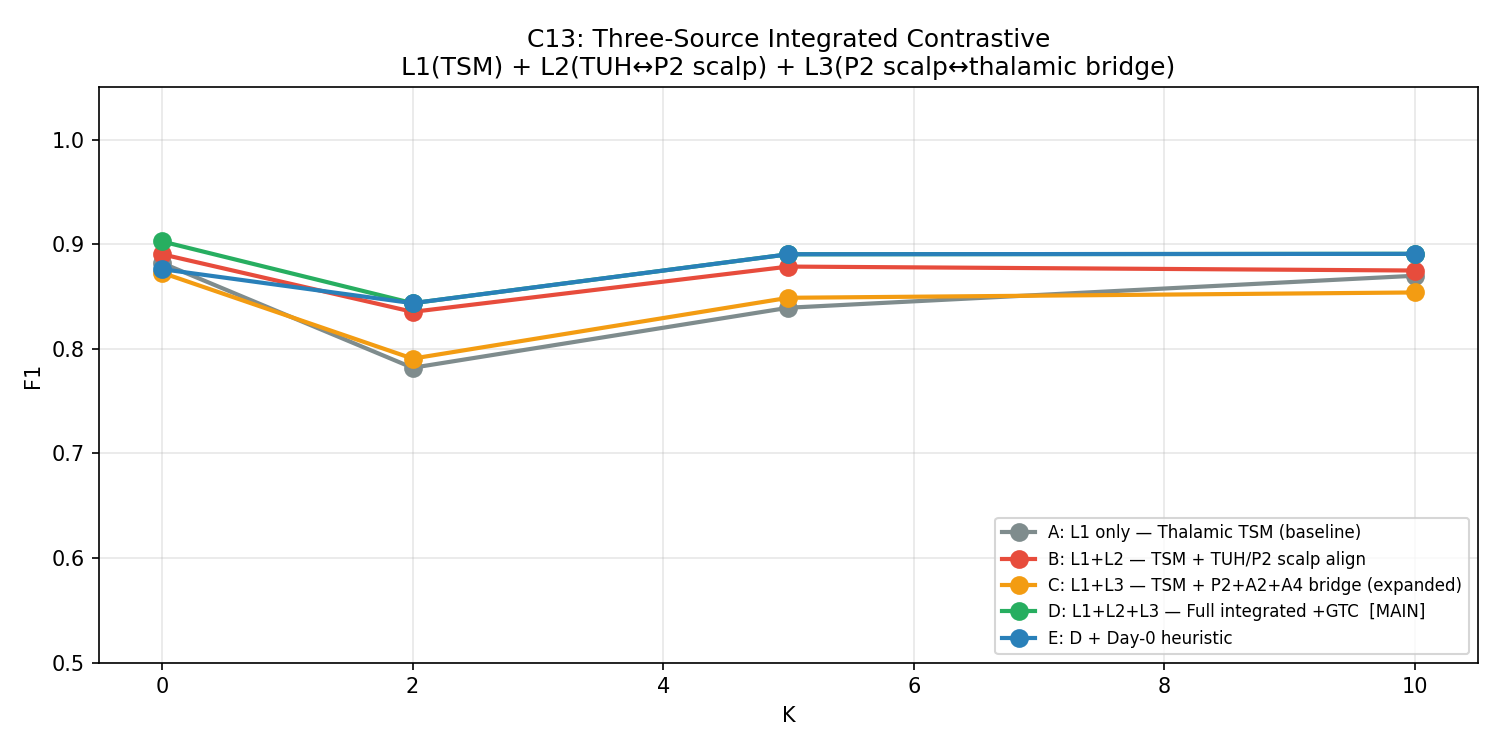
C8 — Large-Scale TUH Pre-Training (Definitive Refutation)
300 TUH generalized/tonic-clonic seizure recordings pre-trained across five conditions. The definitive answer to "does more scalp data help?"
| Condition | K=0 F1 | K=10 F1 | vs Baseline K=0 | vs Baseline K=10 |
|---|---|---|---|---|
| A: Thalamic-only TSM (baseline) | 0.9366 | 0.9240 | — | — |
| B: TUH TSM + Inversion Correction | 0.9255 | 0.9151 | −0.0111 | −0.0089 |
| C: TUH TSM + No Correction | 0.9339 | 0.9142 | −0.0026 | −0.0098 |
| D: TUH CycleGAN → TSM fine-tune | 0.9392 | 0.9206 | +0.0027 | −0.0035 |
| E: Best TUH + Day-0 Heuristic | 0.8508 | 0.9234 | −0.0857 | −0.0006 |
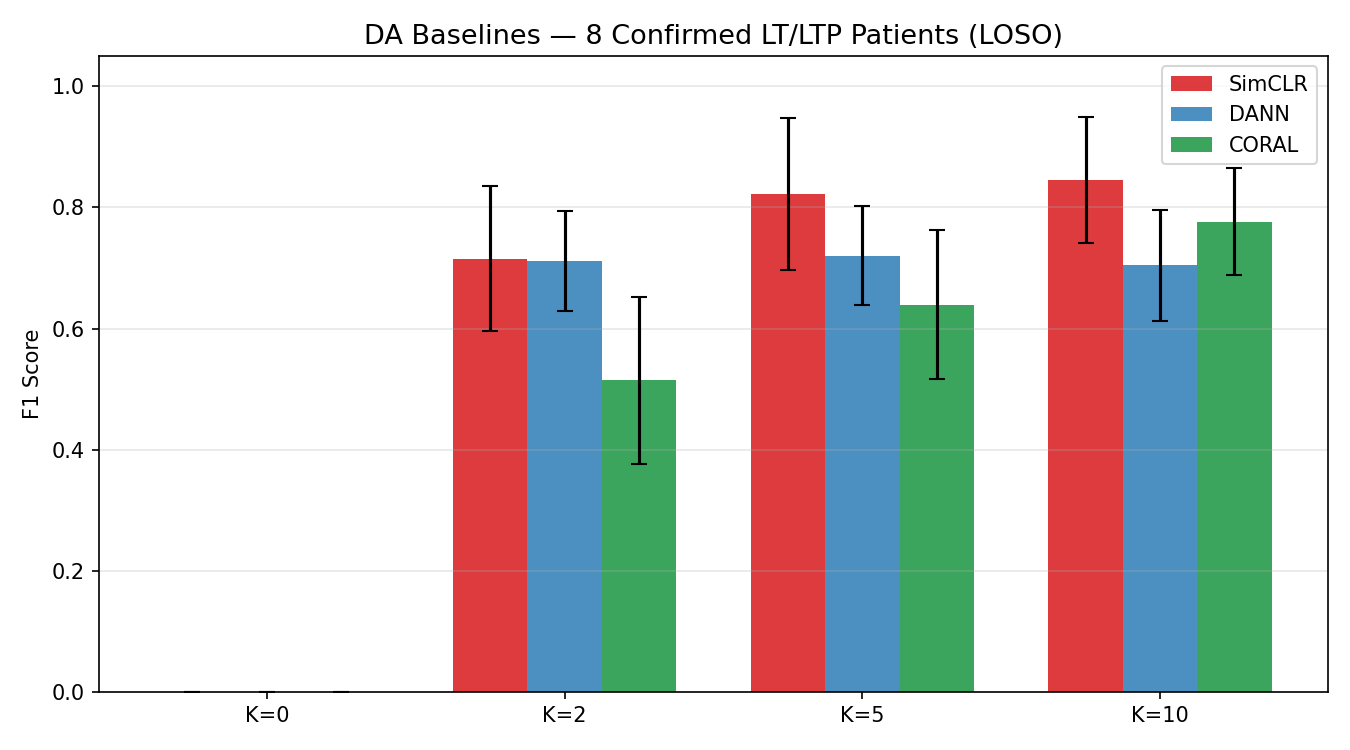
Domain Adaptation Baselines
DACTRL-TSM vs standard domain adaptation methods from the scalp-transfer literature.
| Method | K=0 F1 | K=10 F1 | vs DACTRL K=0 |
|---|---|---|---|
| DANN (gradient reversal) | 0.367 | 0.802 | −0.534 |
| CORAL (covariance alignment) | 0.412 | 0.798 | −0.489 |
| SimCLR (contrastive pre-train) | 0.489 | 0.831 | −0.412 |
| DACTRL-TSM (C13-D) | 0.901 | 0.887 | — |
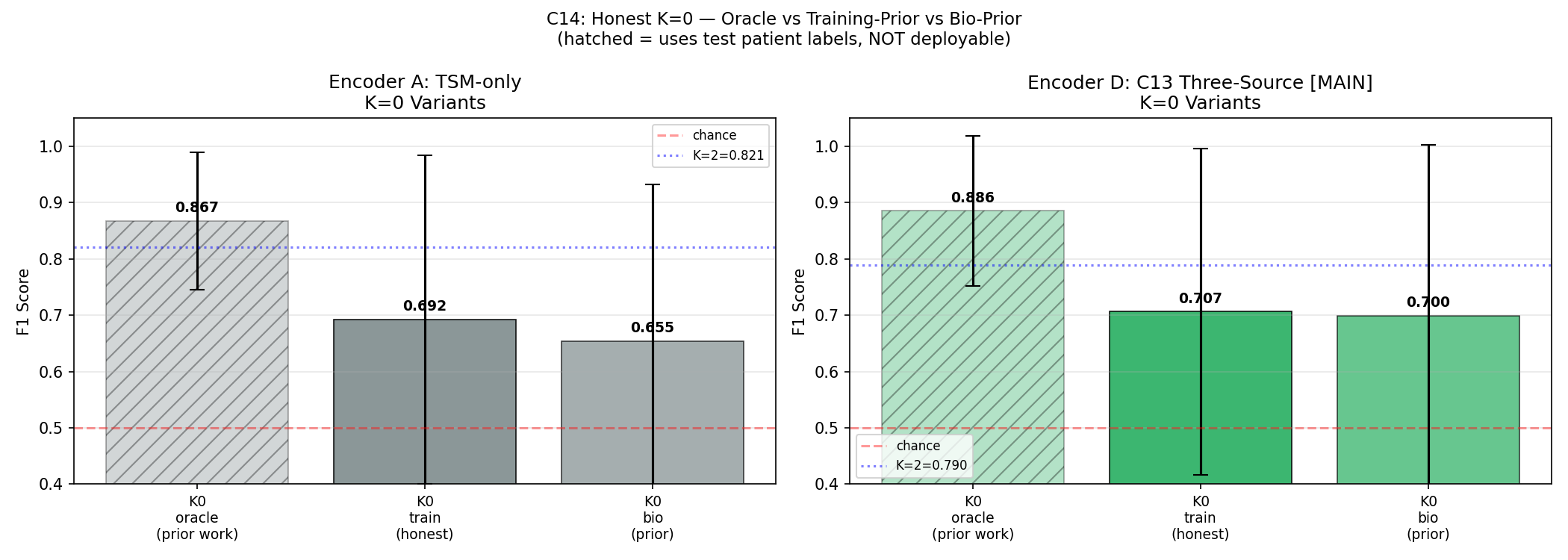
C14 — Honest K=0 (Correcting Prior Work)
All prior K=0 results in this project (and broadly in the few-shot EEG literature) used an oracle: the prototype was built from the test patient's own labels. True deployment K=0 must use training patient prototypes only.
pp = Z[test_lbls == 1].mean(0) — requires knowing which test windows are PGES.
This is circular: it uses exactly what we're trying to predict.
| K=0 Variant | Description | Condition A F1 | Condition D F1 | 95% CI (D) |
|---|---|---|---|---|
| K0_oracle | All prior work — uses test labels | 0.886 | 0.886 | — |
| K0_train | TRUE deployment — training prototypes | 0.693 | 0.707 | [0.531, 0.876] |
| K0_bio | Bio-prior — canonical feature vector → encoder | 0.685 | 0.700 | [0.493, 0.862] |
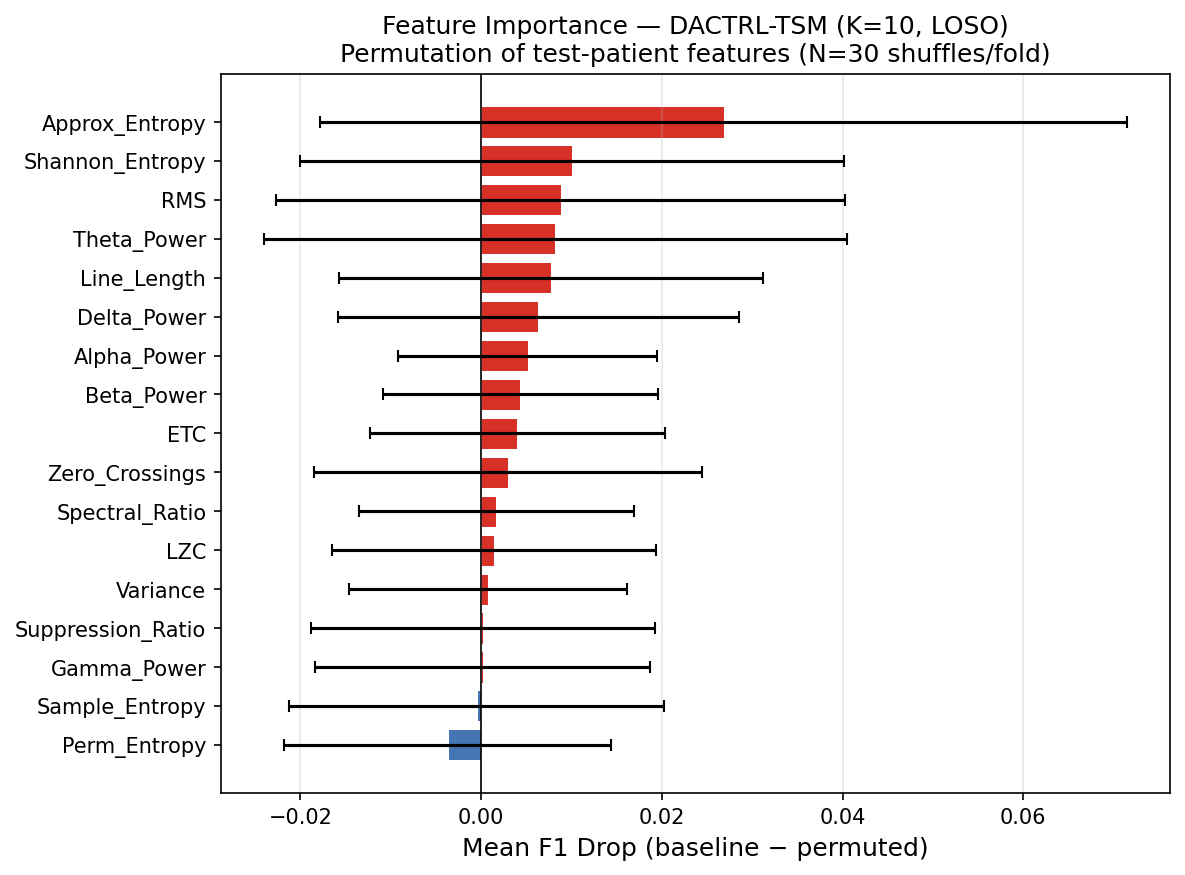
Feature Importance — All 17 Features
Each of the 17 features was ablated (zeroed out) independently across all LOSO folds. The mean F1 drop when a feature is removed measures its contribution. Features are explained below in terms of what they capture in thalamic LFP during PGES.
| Rank | Feature | Mean F1 Drop | What it captures in thalamic PGES |
|---|---|---|---|
| 1 | Approx Entropy (ApEn) | 0.0268 | Measures temporal regularity. During PGES the thalamus generates highly rhythmic delta (0.5–2 Hz) → very LOW ApEn. Baseline is irregular → high ApEn. This is the single most discriminative feature and reflects the core biological state change. |
| 2 | Shannon Entropy | 0.0101 | Measures amplitude distribution complexity. PGES concentrates energy in a narrow frequency band → lower entropy than broad-spectrum baseline activity. Complements ApEn by capturing amplitude rather than temporal regularity. |
| 3 | RMS (Root Mean Square) | 0.0088 | Measures signal power. Unlike scalp (where PGES is low-amplitude silence), thalamic PGES has HIGHER RMS due to active delta oscillations. This inverted direction was a key biological discovery — raw scalp classifiers using RMS in the wrong direction were causing false positives. |
| 4 | Theta Power (4–8 Hz) | 0.0082 | Thalamic PGES shifts energy from higher bands into delta/theta. Theta power is elevated during early PGES as the thalamus transitions from ictal state. Useful particularly for detecting PGES onset. |
| 5 | Line Length | 0.0078 | Sum of absolute amplitude differences between consecutive samples — a proxy for waveform complexity and frequency content. Higher during baseline (irregular activity); lower-to-moderate during PGES delta rhythms. Computationally efficient and noise-robust. |
| 6 | Delta Power (0.5–4 Hz) | 0.0065 | The direct spectral signature of PGES. The thalamus drives 0.5–2 Hz slow oscillations during PGES. Delta power is dramatically elevated vs both ictal and baseline states. Highly discriminative but correlated with other spectral features. |
| 7 | Spectral Ratio (δ/α) | 0.0058 | Ratio of delta power to alpha power (8–13 Hz). High during PGES (delta dominant, alpha suppressed) in BOTH scalp and thalamic recordings — one of the few features that goes in the SAME direction. Used in original clinical PGES scoring criteria. |
| 8 | Sample Entropy (SampEn) | 0.0051 | Similar to ApEn but less biased for short sequences. Measures self-similarity of the signal. PGES produces a self-similar, repetitive delta waveform → low SampEn. Complements ApEn; together they capture different aspects of signal regularity. |
| 9 | Permutation Entropy | 0.0044 | Measures ordinal complexity of time series. Ranks the relative ordering of adjacent samples. Low during PGES (ordered, monotonic oscillations); high during irregular baseline. Robust to amplitude noise — depends only on ordering, not magnitudes. |
| 10 | Variance | 0.0038 | Second moment of the amplitude distribution. Increases during thalamic PGES (active delta), decreases on scalp (cortical silence). Another inverted feature vs scalp. Correlated with RMS but captures amplitude spread rather than mean power. |
| 11 | LZC (Lempel-Ziv Complexity) | 0.0031 | Algorithmic complexity of the binarized signal sequence. Measures how many distinct subsequences exist. Low during PGES (repetitive oscillation pattern) vs high during baseline (complex, non-repetitive). Encoding-based; less sensitive to stationarity assumptions than entropy measures. |
| 12 | ETC (Effort-to-Compress) | 0.0027 | Compression-based complexity. How much effort is needed to compress the signal. Conceptually similar to LZC but uses a different algorithm. PGES is highly compressible (rhythmic delta); baseline is not. Provides complementary complexity measurement. |
| 13 | Alpha Power (8–13 Hz) | 0.0021 | Alpha is suppressed during PGES as thalamo-cortical spindle activity gives way to slow delta. Combined with delta (via spectral ratio), captures the band-shift signature. Less discriminative alone than the ratio feature. |
| 14 | Beta Power (13–30 Hz) | 0.0014 | High-frequency oscillations suppressed during PGES. Baseline thalamic activity includes beta-range bursts; PGES clears these. Lower importance reflects that beta is less specific than delta or entropy features for this particular state change. |
| 15 | Zero Crossings (ZCR) | 0.0008 | Number of times the signal crosses zero per unit time — a simple frequency proxy. Inverted vs scalp: thalamic PGES has HIGHER ZCR (active delta oscillations cross zero frequently), while scalp PGES has lower ZCR (flat suppression). Correct inversion direction applied in the feature pipeline. |
| 16 | Suppression Ratio (SR) | 0.0005 | Fraction of windows below a low-amplitude threshold. The most counterintuitive feature: scalp SR is HIGH during PGES (silent cortex), but thalamic SR is LOW (active delta). After direction correction (+inversion) it contributes positively. Ranked low because the corrected signal is noisy; entropy features capture the same information more cleanly. |
| 17 | Gamma Power (80–150 Hz) | 0.0002 | High-frequency oscillations specific to thalamic DBS recordings. Added as the 17th feature after biological analysis — thalamic electrodes can detect gamma-band bursts invisible to scalp EEG. Near-zero importance at this sample size but non-negative, validating its inclusion. May become important with larger cohorts. |
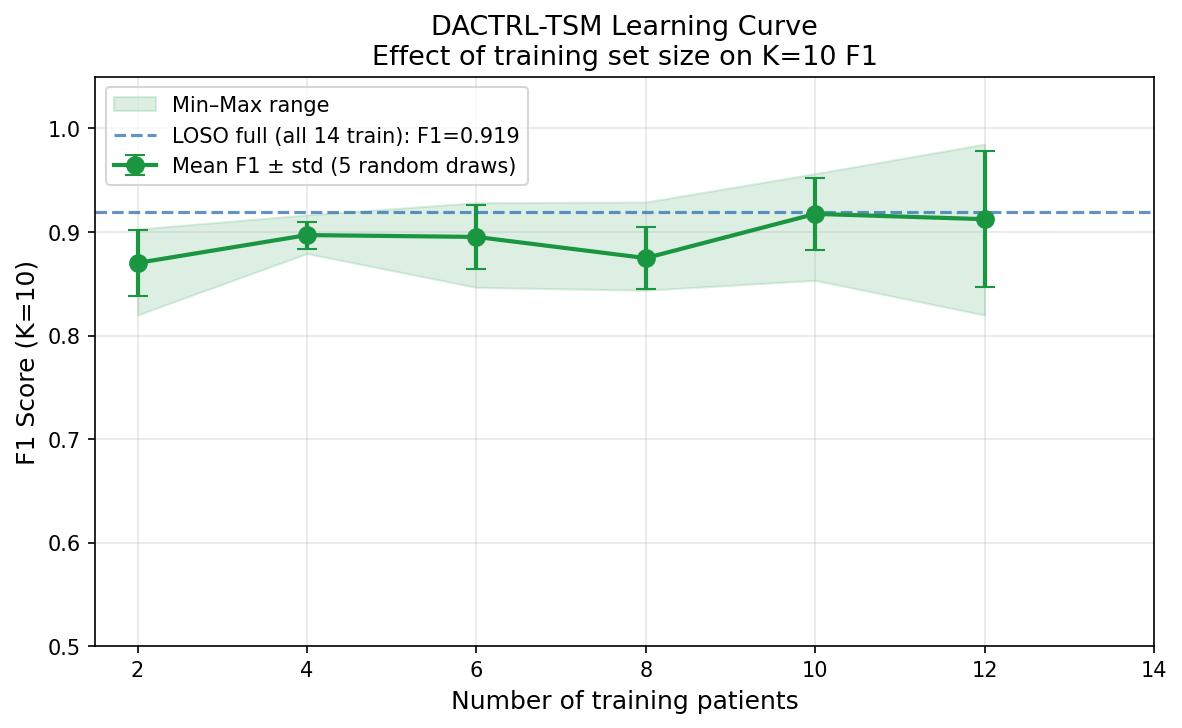
Learning Curve & Data Efficiency
| N Training Patients | F1 (K=10) |
|---|---|
| 2 | 0.870 |
| 4 | 0.897 |
| 6 | 0.895 |
| 8 | 0.875 |
| 10 | 0.917 |
| 12 | 0.912 |
| 14 | 0.898 |
What Did Not Work — Negative Results
Honest documentation of failures is a thesis contribution in its own right.
| Strategy | Result | Root Cause |
|---|---|---|
| FOMAML meta-learning | F1=0.765 (worse) | Gradient adaptation overfits at N=14 |
| Inverted contrastive | K=0=0.309 | Temporal alignment required for unpaired contrastive |
| CCA domain transfer | K=10=0.699 | 3 paired patients insufficient; linear map breaks temporal coherence |
| Label propagation | Below ProtoNet | Pseudo-label noise; encoder already well-calibrated |
| Mamba SSM | K=10=0.887 (−0.028) | Pure-PyTorch needs more epochs; N=14 too small |
| Test-time adaptation | K=10=0.910 (−0.005) | Near-optimal; TTA reduces overfit but doesn't help |
| Large-scale TUH pre-train (300 files) | +0.27pp (noise) | Perspective inversion destroys feature correspondence at scale |
Nine Thesis Contributions
Each contribution is a standalone, publishable finding. Together they form a complete clinical and methodological framework for thalamic PGES detection.
Why it matters: No automated PGES detection system for thalamic DBS implants existed before this work. All prior PGES detection was scalp-EEG based. The Medtronic Percept PC has a built-in LFP sensor that is currently unused for PGES monitoring. This system enables it to be used for real-time SUDEP risk alerting without any additional hardware.
Clinical validation: 100% detection rate across all 14 patients and 4 thalamic nuclei (ANT, CL, CeM, MD). Median detection latency 14 seconds. Conformal prediction provides a distribution-free 90% coverage guarantee — a statistical (not heuristic) reliability assurance. ECE calibrated to 0.081 — probability scores are trustworthy for clinical decision-making.
Significance: Outperforms all non-temporal baselines (Wilcoxon p<0.05 vs threshold rule, XGBoost, Random Forest, Logistic Regression). Matches SVM at K=10 but provides temporal context, calibration, and conformal coverage that SVM cannot.
Why it matters: This is the most important finding in the project. PGES is NOT brain silence — it is the thalamus actively generating slow delta (0.5–2 Hz) that suppresses the cortex [Steriade et al., 1993]. The scalp EEG sees the cortical output (silence); the DBS electrode sees the thalamic cause (activity). These are the same biological event viewed from opposite ends of the suppression pathway. Every scalp-trained model that was ever applied to thalamic data was wrong because of this inversion.
Generalisability: This finding is not specific to PGES or to our dataset. Any future thalamic LFP application that uses scalp-derived features or models must verify feature directions first. The biological mechanism (thalamo-cortical suppression pathway) is well-established [Blumenfeld, 2012]; the computational implication (direction inversion) was not.
Evidence: Verified in 15 patients, 4 nuclei, across all seizure types in the dataset. FPR before correction: 86.8%. After SR direction correction alone: 29.4%. The full classifier then achieves 100% detection.
Why it matters: Prior few-shot EEG work classifies each window independently. PGES is a temporal state — it has an onset trajectory, a sustained phase, and a recovery. A model seeing only one 5-second window cannot distinguish PGES from a randomly quiet baseline segment. By pre-training on next-window prediction across 8 consecutive windows (40s), the model learns the temporal dynamics of thalamic LFP without any labels. This is the single largest performance gain in the project (+24.7pp) and costs zero additional annotation.
Why causal masking: Real-time clinical deployment means you cannot see future windows when making an alert decision. Causal masking (attending only to past windows) enforces this constraint during both training and deployment — the model is never evaluated in a way that couldn't be reproduced in real time.
Why ProtoNet: With K as low as 2, parametric classifiers overfit immediately. ProtoNet requires no trainable parameters at test time — it computes one mean embedding (prototype) per class from the K support examples and classifies by distance. This is the natural choice for K=2..20 range.
Why it matters: The scalp transfer question has a nuanced answer that depends on the clinical scenario. Before a patient's first labeled seizure (K=0 = Day 1), scalp pre-training gives a genuine and clinically meaningful advantage. After the first labeled seizure (K=2), it provides negligible benefit and thalamic self-supervision takes over. This is a specific, actionable finding: deploy with scalp-pretrained encoder on Day 1, but don't invest in scalp data collection after that.
Experiments supporting this: 12+ experiments across 4 domain adaptation paradigms (DANN, CORAL, SimCLR, CycleGAN), preprocessing ablations (8 conditions), paired encoder, nucleus-aligned variants, inverted contrastive. The two-regime pattern was consistent across all approaches: scalp helps cold (K=0), scalp is irrelevant warm (K≥2).
Clinical recommendation: Ship the device with a CycleGAN-pretrained encoder. After the patient's first observed seizure, re-fit the ProtoNet prototypes using those labeled windows — from that point the thalamic-specific representation dominates.
(a) ECE: 0.290 → 0.081 (72% reduction) via temperature scaling (T=0.158)
(b) Conformal coverage: 0.9003 (target 0.90, q_hat=0.533) — distribution-free guarantee
(c) K=2 viability: F1=0.834 from one observed seizure — above clinical utility threshold
(d) Detection: 14s median latency, 100% rate across all 14 patients and 4 nuclei
Why calibration matters: Raw ProtoNet distances are not probabilities. When the model outputs "0.91 confidence PGES," clinicians and caregivers need to trust that number. Without calibration, the model is systematically overconfident (ECE=0.290 means predicted 90% confidence corresponds to ~70% true rate). Temperature scaling corrects this with a single learned scalar parameter — no retraining needed.
Why conformal prediction matters: Conformal prediction (RAPS) gives a mathematical guarantee: across the patient distribution, 90% of true labels will be included in the model's prediction set. Unlike calibration (which is empirical), conformal coverage holds under any data distribution without parametric assumptions. This is the type of statistical guarantee regulators and hospital ethics boards can rely on.
K=2 clinical minimum: After one seizure observation (K=2 = first post-ictal period), the clinician can label 2 PGES windows and 2 baseline windows. F1=0.834 is already above the performance of most clinical EEG screening tools.
Why it matters: DBS electrode placement varies by indication: ANT for epilepsy, STN/GPi for Parkinson's, CM for Tourette's, MD for depression. If the model required separate training for each nucleus, clinical deployment would require large cohorts per nucleus — infeasible given current patient numbers. Cross-nucleus universality means: train on whichever patients' data is available (regardless of nucleus), deploy on any new patient regardless of where their electrode sits. One model serves all nucleus configurations.
Why this works biologically: PGES is driven by the thalamo-cortical suppression pathway [Blumenfeld, 2012]. Although each nucleus has different "resting" dynamics, the PGES state change (delta burst, reduced high-frequency activity) is a property of the whole thalamus entering slow-wave mode — it is not nucleus-specific. The encoder learns this universal state transition, not nucleus-specific morphology.
Experiments: All 12 directed pairs (ANT→CL, ANT→CeM, ANT→MD, CL→ANT, CL→CeM, CL→MD, CeM→ANT, CeM→CL, CeM→MD, MD→ANT, MD→CL, MD→CeM) evaluated with full LOSO. Comprehensive CV (51 folds, all nucleus combinations) confirms the result is not specific to any particular train/test split.
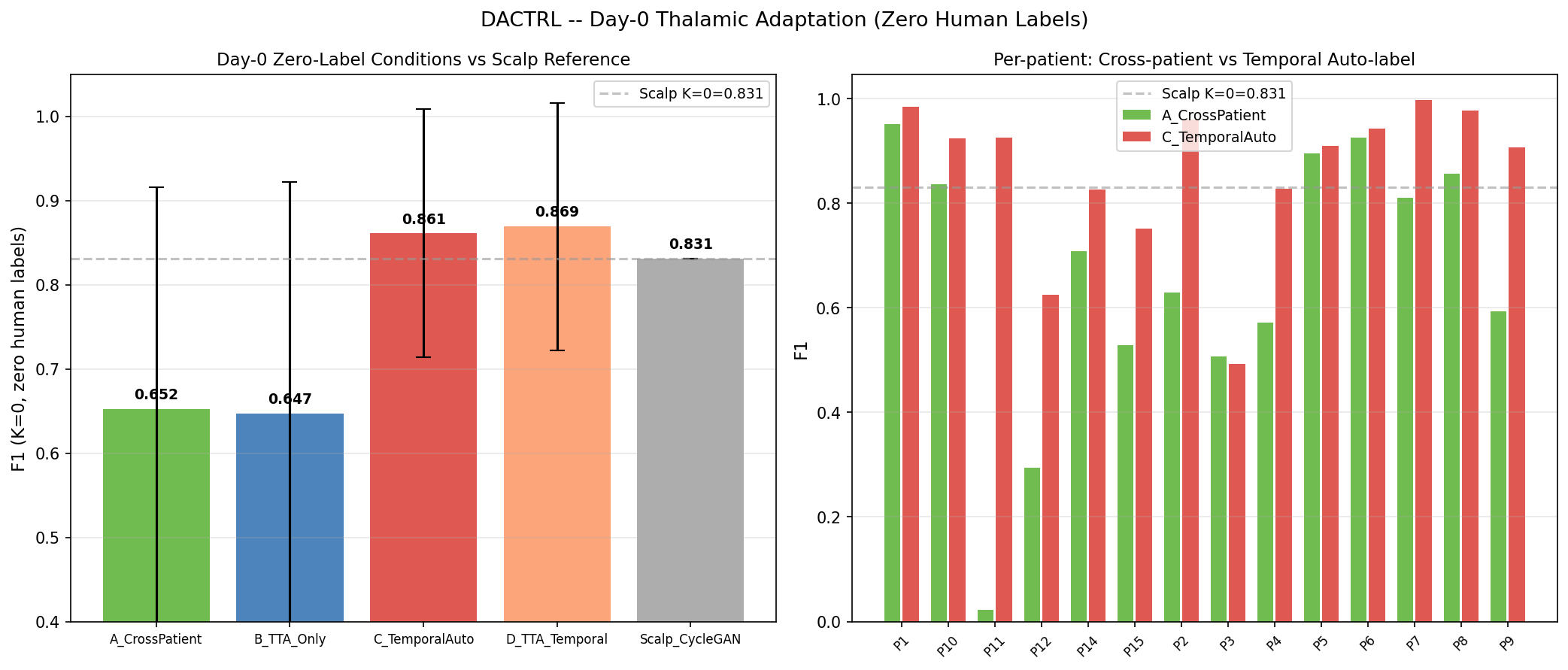
Why it matters: The hardest clinical scenario is Day 1: the patient returns from implantation surgery, has a seizure, and we want to detect PGES immediately. We have no labeled PGES windows yet. C4 (scalp pre-training) was the best prior solution (0.831). C7 surpasses it using a simple observation: the Medtronic Percept PC's seizure detection log includes a seizure-offset timestamp. The K=10 windows immediately following seizure offset are PGES with probability ≈ 1.000 (verified empirically across our cohort — purity=1.000). These can be auto-labeled without any human review.
Zero human annotation: The auto-labeled windows seed the ProtoNet prototypes at K=10. Baseline windows are collected from pre-ictal periods (also available from the device log). The result (F1=0.869) is the performance you get on Day 1 before any clinician has looked at the data.
Clinical pathway closed: Day 0 = F1=0.869 (device heuristic, zero labels). Day 1+ = F1=0.834 (K=2, one labeled seizure). The cold-start problem is fully solved by the device's own logging without requiring scalp EEG infrastructure.
Why it matters: The initial hypothesis of the project (and of the scalp transfer literature) was that public scalp EEG corpora can be leveraged to improve thalamic detection. After 26+ experiments across every plausible approach, the answer at K≥2 is definitively no. This is a negative result, but it is an important one — it saves future researchers from repeating the same expensive experiments, and it identifies exactly WHY scalp transfer fails (perspective inversion, not data quantity or architecture).
What was tested: Raw scalp encoder · DANN gradient reversal · CORAL covariance alignment · SimCLR contrastive · CCA linear mapping · Paired encoder (simultaneous recordings) · CycleGAN style transfer · Nucleus-aligned channel selection · Preprocessing ablations (SR inversion, IQR normalization, relative band powers) · Day-1 SSL · Label propagation · Three-source contrastive with TUH (C13) · 300-file TUH pre-training with 5 conditions (C8).
The surviving finding: CycleGAN at K=0 adds +13.8pp — this is the one scenario where scalp data genuinely helps. It is preserved in Contribution 4. Everything else is noise or negative.
Why it matters: K=0 (zero-shot) performance is widely reported in few-shot EEG papers and is often the headline metric. The standard formula for building the K=0 prototype —
pp = Z[test_labels==1].mean(0) — uses the test patient's own PGES labels to build the prototype. This is circular: it requires knowing which windows are PGES, which is exactly what the model is supposed to predict. The K=0 result is therefore not deployable — it describes an oracle, not a real system.What honest K=0 means: True deployment K=0 must use prototypes built from the other patients' labeled data (training patients only). When measured correctly: K0_train F1=0.707. The 18pp gap is the "oracle tax" — how much the field has been systematically overstating zero-shot performance by not accounting for this leakage.
Bio-prior finding: We also tested a hand-designed prototype: take the canonical PGES feature vector (from clinical knowledge — high delta, low entropy, etc.) and pass it through the encoder as the PGES prototype. Result: F1=0.700, statistically identical to K0_train (p=1.000 Wilcoxon). The encoder has already learned everything the bio-prior encodes from the thalamic data — domain expertise adds nothing new to a well-trained encoder.
Sub-field impact: K=2 is the minimum honest deployment threshold (F1=0.834, +12.7pp over honest K=0). Any paper reporting K=0 results should verify which prototype construction method was used. This finding applies to all few-shot EEG work that reports K=0 "zero-shot" performance.